Table Filtering
Submissions





GlyTouCan
Glycan Structure Repository
GlyComb
Glycoconjugate Repository
GlycoPOST
Glycomics MS raw data RepositoryUniCarb-DR
Glycomics MS Repository for glycan annotations from GlycoWorkbenchAll Resources
Genes / Proteins / Lipids Glycans / Glycoconjugates Glycomes Pathways / Interactions / Diseases / Organisms GlycoBase
GlycoBase
GlycoBase is a relational database which contains the HPLC elution positions for over 350 2-AB labelled N-glycan structures together with predicted products of exoglycosidase digestions.
GlycoBase and autoGU: tools for HPLC-based glycan analysis
Campbell MP, et al.
Bioinformatics. 2008 May 1;24(9):1214-6. PMID: 18344517
| Source | Last Updated |
|---|---|
| Glycobase | April 18, 2024 |
| GlyTouCan ID | GlycoBase ID ▲ | Compositions | Compositions (GlyTouCan ID) | Mass | GU (average) | GU (standard deviation) |
|---|---|---|---|---|---|---|
| 942 | Fuc2HexNAc6Hexose7NeuAc4 |
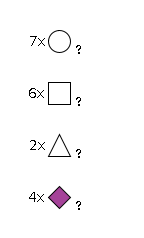 G71560PC
G71560PC
|
3827.35 | 11.9 | 11.9 | |
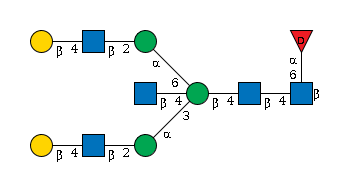 G53000RQ
G53000RQ
|
943 | Fuc1HexNAc5Hexose5 |
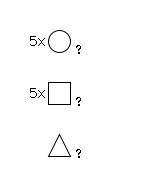 G91636VS
G91636VS
|
1989.73 | 8.86111 | 1107.64 |
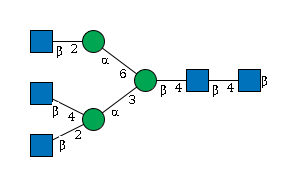 G29011JC
G29011JC
|
944 | HexNAc5Hexose3 |
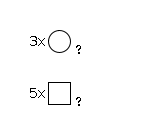 G31936TA
G31936TA
|
1519.57 | 6.23547 | 586.135 |
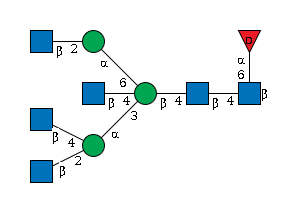 G75355YV
G75355YV
|
945 | Fuc1HexNAc6Hexose3 |
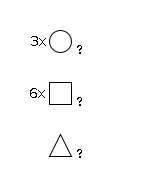 G50757KG
G50757KG
|
1868.7 | 6.38286 | 172.337 |
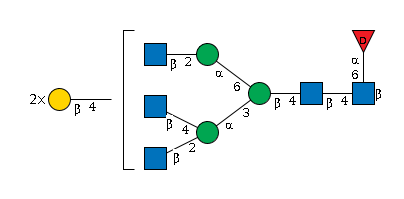 G17580TM
G17580TM
|
946 | Fuc1HexNAc5Hexose5 |
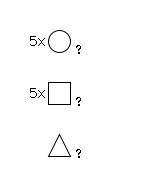 G91636VS
G91636VS
|
1989.73 | 7.77783 | 38.8895 |
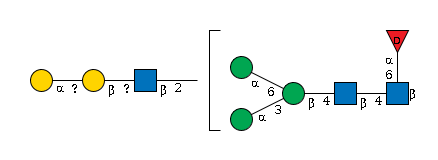 G53770LT
G53770LT
|
947 | Fuc1HexNAc3Hexose5 |
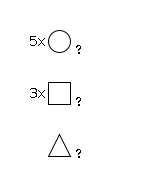 G82119TF
G82119TF
|
1583.57 | 7.08 | 0.0 |
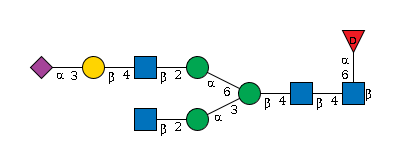 G70069EN
G70069EN
|
948 | Fuc1HexNAc4Hexose4NeuAc1 |
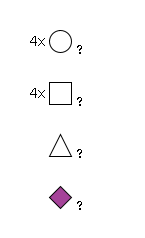 G14994KB
G14994KB
|
1915.69 | 7.79429 | 101.326 |
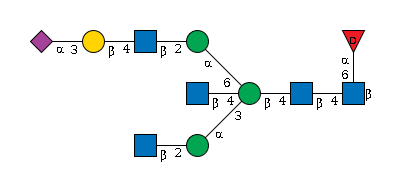 G63075AF
G63075AF
|
949 | Fuc1HexNAc5Hexose4NeuAc1 |
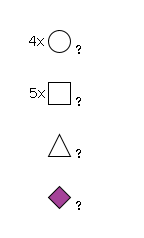 G72667IM
G72667IM
|
2118.77 | 7.94083 | 87.3494 |
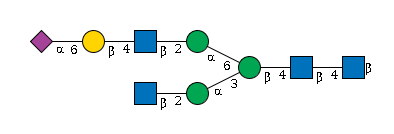 G90725ZC
G90725ZC
|
950 | HexNAc4Hexose4NeuAc1 |
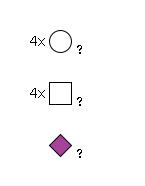 G31916IQ
G31916IQ
|
1769.63 | 5.76529 | 92.2487 |
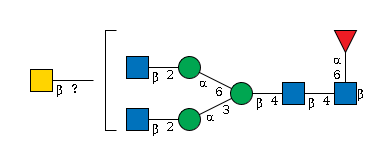 G74120KH
G74120KH
|
951 | Fuc1HexNAc5Hexose3 |
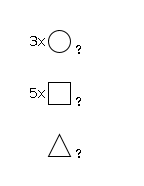 G10256JP
G10256JP
|
1665.62 | 6.39 | 0.0 |
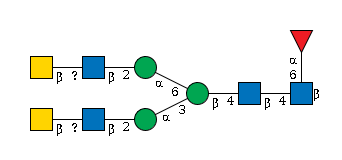 G22562DG
G22562DG
|
952 | Fuc1HexNAc6Hexose3 |
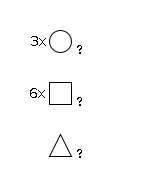 G50757KG
G50757KG
|
1868.7 | 6.92 | 0.0 |
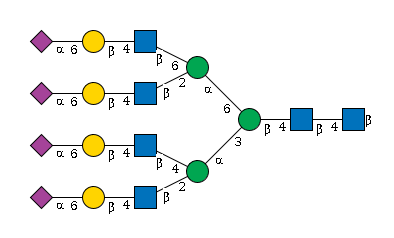 G15998WM
G15998WM
|
954 | HexNAc6Hexose7NeuAc4 |
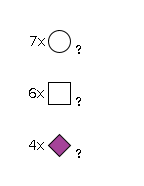 G66088HZ
G66088HZ
|
3535.24 | 11.29 | 22.58 |
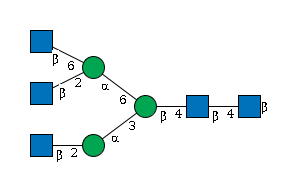 G10219AA
G10219AA
|
955 | HexNAc5Hexose3 |
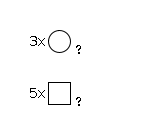 G31936TA
G31936TA
|
1519.57 | 6.05933 | 84.8311 |
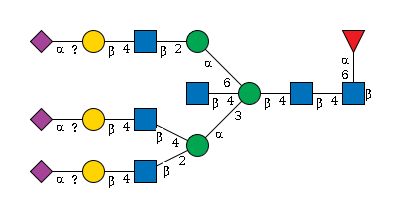 G76297WR
G76297WR
|
956 | Fuc1HexNAc6Hexose6NeuAc3 |
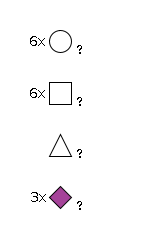 G62461SM
G62461SM
|
3228.15 | 10.49 | 31.4707 |
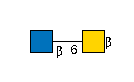 G23447MZ
G23447MZ
|
9572 | HexNAc2 |
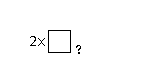 G71838YU
G71838YU
|
424.169 | 1.795 | 1.79501 |
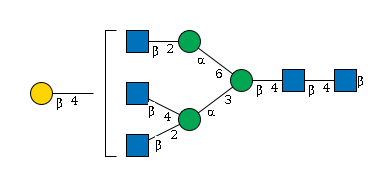 G90426CG
G90426CG
|
963 | HexNAc5Hexose4 |
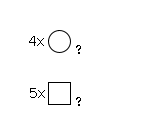 G23432EQ
G23432EQ
|
1681.62 | 6.7 | 0.0 |
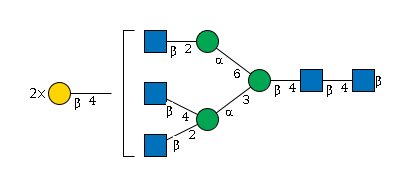 G00670OT
G00670OT
|
964 | HexNAc5Hexose5 |
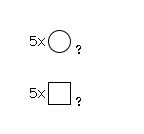 G79568CQ
G79568CQ
|
1843.67 | 7.52 | 7.52003 |
 G36107HL
G36107HL
|
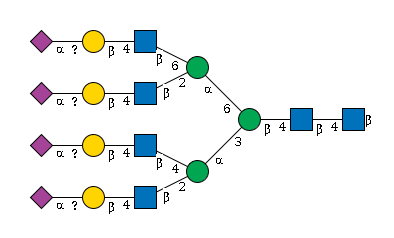
965 | HexNAc6Hexose7NeuAc4 |
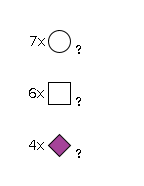 G66088HZ
G66088HZ
|
3535.24 | 10.88 | 32.64 |
 G24720OO
G24720OO
|
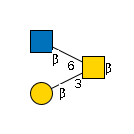
9688 | HexNAc2Hexose1 |
 G32550BI
G32550BI
|
586.222 | 2.71833 | 13.5917 |
